Protein detection is a fundamental and critical technology in biochemistry, molecular biology, and medical diagnosis. There are usually many methods for detecting proteins based on different purposes, such as qualitative, quantitative, localization, and functional analysis. This article introduces several commonly used protein detection methods to familiarize you with protein detection techniques.
What is protein?
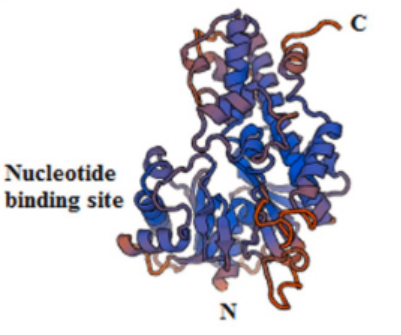
Proteins are substances with a certain spatial structure formed by the twisting and folding of polypeptide chains composed of amino acids through dehydration condensation. It is an important substance that constitutes living organisms.
The properties of proteins
• Physicochemical properties
Proteins are amphiphilic substances, with amino (N-terminal) and carboxyl (C-terminal) groups present in their molecules.
• salting out
Adding high concentration neutral salt solution can reduce the solubility of proteins, causing them to precipitate without changing their original properties.
• sex reassignment
Proteins undergo irreversible changes and lose their original function and activity under the action of heat, acid, alkali, UV, and heavy metals.
Common detection methods for proteins
01,Used to determine the total concentration of proteins in a solution, it is fast and simple, but usually cannot distinguish between different types of proteins.
| detection method | principle | advantage | shortcoming | Applicable scenarios |
| UV absorption method | The tryptophan and tyrosine residues in proteins exhibit strong absorption peaks at 280nm. The concentration can be estimated using standard curves or empirical formulas (1 OD280 ≈ 1 mg/mL). | Fast, non-destructive, sample recyclable | Greatly influenced by protein specific amino acid composition; Nucleic acid also absorbs at 280nm, which can interfere with the measurement. | Preliminary and rapid estimation of the concentration of relatively pure protein samples. Real time monitoring of elution peaks during chromatography. |
| Bradford Law | Coomassie Brilliant Blue binds to the hydrophobic region of proteins under acidic conditions, changing from reddish brown to blue. The absorbance is measured at 595nm | Fast, sensitive, and with minimal interference | Fast, sensitive, and with minimal interference | Rapid quantification of conventional cell lysate and relatively pure protein samples. Laboratory daily methods |
| Lowry method | In two steps: the protein reacts with copper ions in alkaline solution (biuret reaction) to form a copper protein complex, which reduces phosphomolybdic acid phosphotungstic acid reagent (Folin phenol reagent) to produce a deep blue color. | Sensitivity higher than UV method | The steps are cumbersome; Severe interference from various substances such as Tris, sucrose, EDTA, etc | Currently, it is rarely used, mainly in historical methods or specific non-interference systems |
| BCA method | Similar to the Lowry method, proteins reduce Cu ² ⁺ to Cu ⁺ under alkaline conditions. BCA reagent binds to Cu ⁺ to form a purple complex, which is detected at 562nm | High sensitivity, strong anti-interference ability (good compatibility with detergents), easy to operate | The response of different proteins still varies | Accurate quantification of complex samples containing detergents, such as cell and tissue lysates. Scenarios that require high sensitivity |
02,The qualitative and quantitative analysis of specific proteins can detect specific target proteins in a mixture.
| detection method | principle | advantage | shortcoming | Applicable scenarios |
| SDS-PAGE (sodium dodecyl sulfate polyacrylamide gel electrophoresis) | SDS uniformly charges proteins with negative charges, and the migration rate during electrophoresis depends only on the molecular weight. | Strong separation ability: Separate mixtures by molecular weight. Qualitatively evaluate purity and molecular weight. low cost | Only semi quantitative (staining intensity) is possible. Cannot identify proteins. Low flux | Analyze protein sample purity and estimate molecular weight. The pre steps for Western Blot and mass spectrometry analysis. |
| Enzyme linked immunosorbent assay (ELISA) | Fix antigens or antibodies on a solid surface, react with enzyme labeled antibodies, add substrates, and develop color. The color depth is proportional to the amount of target protein. | High specificity and sensitivity. It can be absolutely quantified (with a standard curve). High throughput: suitable for processing large amounts of samples (96/384 well plates). The operation is relatively standardized. | High quality antibodies that require pairing (sandwich method). Unable to provide molecular weight information. | High throughput and precise quantification of specific proteins in body fluids (serum, culture supernatant). Core technologies for clinical diagnosis and drug development. |
| Western Blot (protein immunoblotting) | Separate the samples by molecular weight through SDS-PAGE electrophoresis, then transfer them onto membranes (such as PVDF, NC membranes) and incubate and color them with specific antibodies | High specificity: Antibodies determine specificity. Provide multiple pieces of information: presence, molecular weight (post-translational modifications), semi quantitative. High sensitivity (Pick level) | The operation is complex and time-consuming (1-2 days). The quantitative linear range is narrow and semi quantitative. Highly dependent on antibody quality (specificity, potency). Low flux. | Confirm the presence, molecular weight changes (such as cleavage, glycosylation), and relative abundance of specific target proteins in complex samples. The gold standard in scientific research |
| Immunoprecipitation and immunoprecipi- tation(IP/Co IP) | Immunoprecipitation and immunoprecipi- tation (IP/Co IP) | Can enrich target proteins from complex mixtures. Can study protein interactions (Co IP). Flexible downstream analysis (WB, MS) | Non physiological conditions: May capture false positive interactions. The background may be higher. Low efficiency in capturing weak interactions. | Enrichment of low abundance proteins, validation or discovery of protein-protein interactions |
03,Used for precise quality determination such as protein identification and structural analysis.
| detection method | principle | advantage | shortcoming | Applicable scenarios |
| Mass spectrometry (MS) | Enzymatically hydrolyze proteins into peptide segments and separate them based on their hydrophobicity, charge, size, and other characteristics | Gold standard: Can accurately identify unknown proteins. Ultra high resolution capability: capable of analyzing post-translational modifications (phosphorylation, glycosylation, etc.). High throughput: One experiment can identify thousands of proteins | The instrument is extremely expensive and the operation is complex. High pre-processing requirements. Quantitative complexity and limited dynamic range (high abundance proteins can easily mask low abundance proteins). Cannot directly analyze activity | Proteomics, identification of unknown proteins, analysis of post-translational modification profiles, and discovery of biomarkers. |
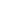
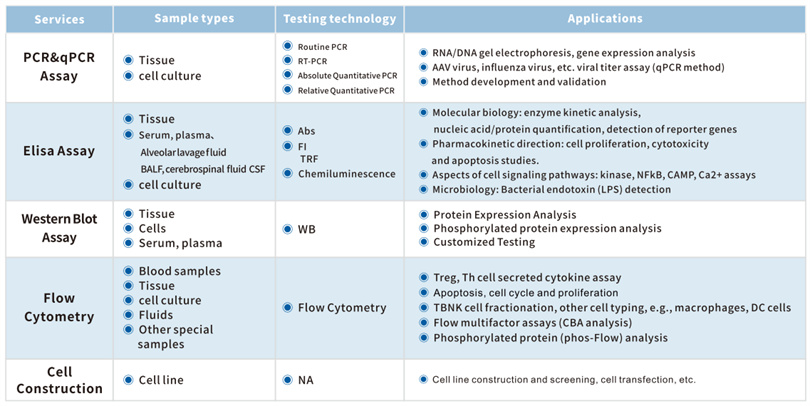
The KCI • KMQ molecular biology platform has advanced technological means and rich project experience. It can not only undertake biochemical detection indicators with a daily detection volume of over 300, but also provide customers with various molecular biology detection methods including flow cytometry, qPCR, ELISA, WB, IP/Co IP, etc. At present, the company has established extensive long-term cooperation with many well-known pharmaceutical companies and research institutions at home and abroad, providing a solid foundation for the development of innovative drugs.